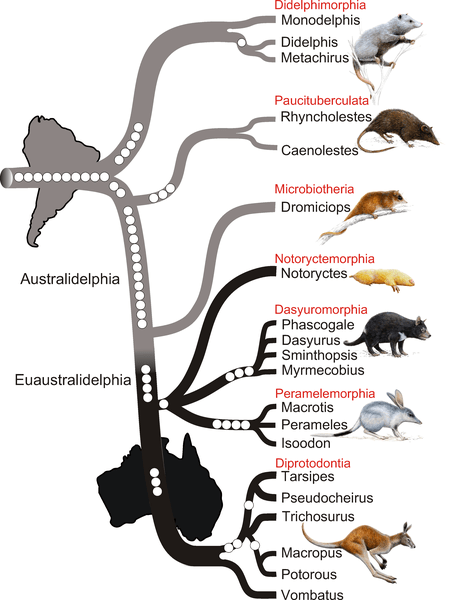
The phylogenetic relationships between two orders of marsupials have been intesively debated. Authors benefited from recent sequencing projects which provided two marsupial genomes: this of the South American opossum (Monodelphis domestica)and the one of a kangaroo, the Australian tammar wallaby (Macropus eugenii).
Retroposons are suitable and homoplasy-free markers: their insertion sites are random; parallel insertions or exact excisions are very rare. Thus, if one finds a retroposon in the homologous genomic loci of both species this indicates a common ancestry; on the contrary: if the marker is missing in one of the species, it means prior divergence. Moreover, one retroposon can insert into another (i.e. transposition into transposition). These nested mobile elements insertions provide precious information about the relative times during which given retroposon families integrated into genomes: young elements can insert into older ones, but the reciprocal is impossible.
The results in a nutshell
After the complete screening of the opossum and kangaroo genomes, authors found ~8,000 and ~4,000 nested retroposon insertions, respectively. Then, the frequencies and time scales of SINEs (Short INterspersed Elements) were calculated (using TinT software). Three SINE groups are thus identified:
- Those specific to the lineage leading to opossum => phylogenetically informative markers present in the opossum lineage;
- These specific to the lineage leading to kangaroo => phylogenetically informative markers present in the kangaroo lineage;
- Those active in both species => phylogenetically informative markers present in both lineages.
Also, three different strategies enabled the authors to identify ~220,000 genomic loci containing retroposons. After screening and experimental confirmation, the alignment and analysis of a total of ~440 marsupial sequences revealed 53 informative markers. Ten of those confirmed again the monophyly of marsupials. The other 43 phylogenetically informative retroposon markers provide significant support for most of the basal splits within marsupials.
Authors did not find any loci containing elements present in opossum plus Paucituberculata but absent in kangaroo, which would have supported the alternative of a close relationship between Didelphimorphia and Paucituberculata. They screened for markers that would support the alternative hypothesis of Paucituberculata being the sister to all marsupials. Experimental verification showed that all of the putative elements were also present in the order Paucituberculata (Rhyncholestes), thus supporting the monophyly of marsupials, but not the basal divergence.
Furthermore, 13 of the original 53 markers were present in the South American Microbiotheria and the four Australasian orders, but not in either Didelphimorphia or Paucituberculata. This observation significantly supports the monophyly of Australidelphia. The branch separating Australidelphia from Didelphimorphia and Paucituberculata is one of the strongest supported as well. Nevertheless, the poor fossil record from South America, Antarctica, and Australia does not allow to assess Australidelphian early relationships and biogeography.
Discussion
Two competing hypotheses exist regarding Microbiotheria. The latter are either excluded from the Australasian order (on the basis of nuclear protein-coding genes); or embedded into it (completely or partially based on mitochondrial data). No reliable marsupial phylogeny exists today. In the present study, authors provide evidence for four independent diagnostic retroposon insertions. The latter allow placing Microbiotheria within South America marsupials.
Thus, the authors propose the new name Euaustralidelphia. It would encompass the monophyletic grouping of four Australasian orders: Notoryctemorphia, Dasyuromorphia, Peramelemorphia, and Diprotodontia. In total, 18 out of the initial 53 retroposon markers provide significant support for the monophyly of each of the five multi-species marsupial orders.
Authors conclude:
“the retroposon marker system identified a clear separation between the South American and Australasian marsupials. Thus, the current findings support a simple paleobiogeographic hypothesis, indicating only a single effective migration from South America to Australia, which is remarkable given that South America, Antarctica, and Australia were connected in the South Gondwanan continent for a considerable time.”
![]()
Nilsson MA, Churakov G, Sommer M, Tran NV, Zemann A, Brosius J, & Schmitz J (2010). Tracking marsupial evolution using archaic genomic retroposon insertions. PLoS biology, 8 (7) PMID: 20668664